|
NSCL DDAS
1.0
Support for XIA DDAS at the NSCL
|
Every experiment is different and produces data in different format. For this reason, there is no precompiled version of SpecTcl provided to deal with DDAS data. Rather there are two unpackers that are provided to use, DAQ::DDAS::DDASUnpacker and DAQ::DDAS::DDASBuiltUnpacker. The difference between the two is that the latter unpacks data that has been built with the event builder. The former unpacks data as it would be outputted from the Readout program. Both provide consistent interfaces for interacting with the DDAS data. In this section, you will learn how to incorporate these into a SpecTcl. The source code of the final product is available installed at /usr/opt/ddas/VERSION/share/src/multicrate_src on NSCL spdaq machines. For other machines that choose to install to a different prefix, it is installed at @prefix@/share/src/multicrate_src.
The user does not have deal with the low-level raw data when building a SpecTcl. Rather, they just need to implement a class that uses the unpacked DDAS data to set TreeParameters for histogramming. In this way, the user from the details of the DDAS data structure. In fact, you should never need to write your own DDAS event parser. We provide tools for this for SpecTcl and other raw data. In any case, let's get down to business constructing a tailored SpecTcl.
First you need to copy the SpecTcl skeleton files into a directory where you will build your tailored SpecTcl. You can find the skeleton files at /usr/opt/spectcl/VERSION/Skel. This would be done with something like:
mkdir myDevelopmentDir cp /usr/opt/spectcl/3.4-005/Skel/* .
Any SpecTcl version newer than 3.4 will be suitable.
The first thing that you need to do is decide how we want to structure our TreeParameters. TreeParameters are the typical mechanism that a user will use to update a spectrum. For each event, if the TreeParameter has been assigned a value, it will be histogrammed unless the event is marked as bad. Typically, experimenters like to structure their TreeParameters in a tree-like structure. In this tutorial, we will do similar. We are going to concern ourselves with the energy and timestamp values for each channel as well as how many channels had data in each event (i.e. multiplicity). Because there are three modules in our setup and each has sixteen channels, there are 48 channels we need to allocate TreeParameters for. I will layout the structure of the data in a tree structure where there is a top-level event structure called MyParameters that holds the multiplicity information as well as all of the channel data object. The code in the header file is:
The implementation file is very simple:
Now that we have laid out the structure of the TreeParameters we will turn our attention to assigning data to them. Normally, users would be required to parse raw data in their SpecTcl and then assign values to the tree parameters. In the DDAS support, we parse the raw data for you. We will name the header file for our mapper MyParameterMapper.h. It has the following contents:
Note that there is not much to this class. The class maintains a reference to our parameters and an m_chanMap data member, which we will talk about a little later. As far as methods, there is a constructor, the mapToParameters(), and computeGlobalIndex() methods. The guts of the logic for the class is in the mapToParameters method.
So what does the mapToParameters() method do? Well, let's consider what is passed into the method as arguments. The most important of these is the first, which is a vector (vector = resizable array) of DDASHit objects. A DDASHit object encapsulates the data contained in a single channel event. It has things like the energy, timestamp, raw data elements, geographic information, trace data, as well as QDC and energy sum / baseline data. It is essentially just a vehicle to access data elements at a higher level. The provided unpacker will parse the raw data and fill the vector with all of the data in each event. If more than one DDAS hit was in an event, there will be more than one DDASHit object in the vector.
Here is the implementation of the mapToParameters() method:
Let's think a little about the computeGlobalIndex() method. The responsibility of this method is to map the hit to a global channel index, index in range [0, 47], using the crate, module, and channel information. To do this, we need to input some information concerning the layout of the modules in the crates. What is most beneficial is to know the global index of the first channel of the first module in each crate. Now I set the crate id of my first crate as 0 and the second crate as 2. That is where the m_chanMap comes in. The first crate has 2 modules in it, so the first channel of the second crate will begin at 32. See how this is defined in the constructor:
The mapToChannels method then uses this m_chanMap in the following way:
That is it for defining our parameters and mapping. We will now turn to incorporating this into SpecTcl.
The MySpecTclApp.cpp and Makefiles both need to be modified. First we will consider the MySpecTclApp.cpp. In MySpecTclApp.cpp, locate the MySpecTclApp::CreateAnalysisPipeline() method, which will have an implementation already. We will replace the implementation with a simpler one that look like this:
We will also redefine the definition of "Stage1" in the global scope. Locate where it says:
Replace this with
To round out our work on MySpecTclApp.cpp, you simply need to add some include directives to the top of the file. These will bring the MyParameters, MyParameterMapper, DAQ::DDAS::CDDASBuiltUnpacker classes into scope. Add these lines to the top:
The final work that needs to be done is to modify the Makefile. Add the MyParameters and MyParameterMapper class to the build by adding MyParameters.o and MyParameterMapper.o to the OBJECTS variable. It should look like this when you are done:
Next the compiler needs to be told where to find the DDASBuiltUnpacker.h file and that we need c++11 mode. That is done by adding to the USERCXXFLAGS so it looks like this:
Finally, we need to tell the linker where the precompiled DDAS code is. Add to USERLDFLAGS so it looks like this:
where VERSION is the same as in the very beginning of this tutorial.
With those changes, you should be able to build your SpecTcl. This is accomplished by typing:
make
To run the compiled SpecTcl, execute the command:
./SpecTcl
To remind you, the source code for the example can be found at /usr/opt/ddas/VERSION/share/src/multicrate_src.
The simplest way to create spectra is to run your tailored SpecTcl and use its user interface to create an array of spectra for the parameters we created.
Here's the SpecTcl treeparameter user interface after we start our SpecTcl:
module-1 in the spectrum name entry to set the base spectrum name.File->Save... menu command to save these definitions.You can now attach SpecTcl to the online system using the Data Source->Online... Menu entry. In the resulting dialog:
Start a run. Once data starts making its way out of the event builder, you should be able to see counts in the histograms that have valid inputs. Here's a sample pulser spectrum from my tests:
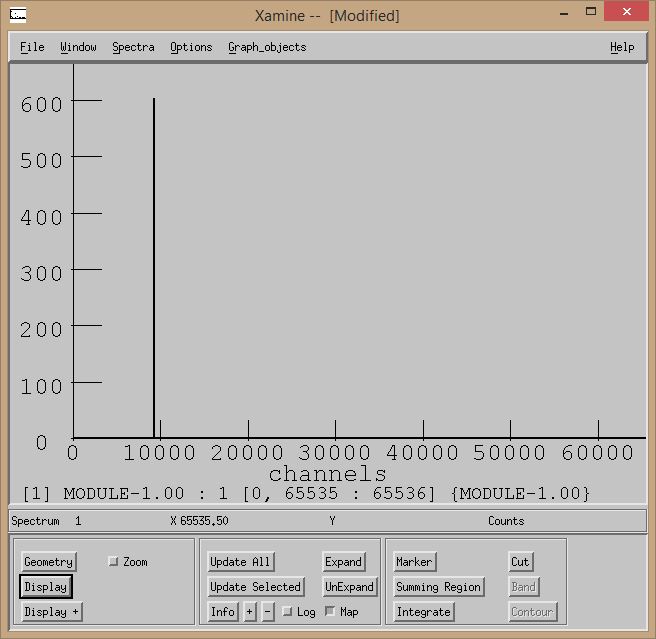
 1.8.1.2
1.8.1.2